Coot: Difference between revisions
No edit summary |
|||
| Line 73: | Line 73: | ||
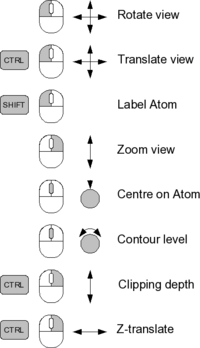
[[Image:Coot-controls-small.png|200px]] | [[Image:Coot-controls-small.png|200px]] | ||
===Stereographic Display === | |||
==== Side-by-Side ==== | |||
==== Hardware Stereo ==== | |||
==== Zalman Stereo ==== | |||
===External Links=== | ===External Links=== | ||
Revision as of 07:11, 20 August 2009
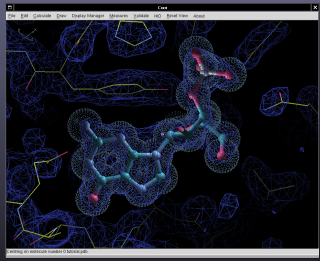
Coot is a graphics program for building, refining and analysing macromolecular models obtained with crystallographic procedures.
There is a homepage with extensive documentation. The program may be downloaded for Linux, Mac and Windows computers from the primary server (as of July 10, 2009 there's a new site at http://www.biop.ox.ac.uk/coot/devel/build-info.html) or, if that is not available, from an external mirror. The license of Coot is GNU GPL.
Installing Coot
Installing Coot on OS X
Please refer to the Installing Coot on OS X page
Installing Coot on Windows
Please refer to the WinCoot install and download page.
Installing Coot on Linux
Installing coot on linux is rather more straightforward than on OS X, because most linux systems are based on gnome and/or kde, and tend to have many of the required components already installed. Most of the other dependencies are also readily available.
Installation from a distributed binary tarball package
This is the recommended way for those who do not want to delve into the mysteries of compiling and linking a great but complex piece of software. Read the (somewhat outdated, it seems) Coot FAQ to find "Additional Notes" for your operating system.
In short, just go to http://www.ysbl.york.ac.uk/~emsley/software/binaries/nightlies/pre-release/ and pick a suitable binary, e.g. coot-0.5-pre-1-revision-1003-binary-Linux-i386-fedora-5.tar.gz for a Red Hat Enterprise Linux 5 or CentOS-5 system (Fedora 6 corresponds to RHEL5, thus Fedora 5 binaries are OK). If you prefer a "stable" binary, these are at http://www.ysbl.york.ac.uk/~emsley/software/binaries/stable/.
Then un-tar it under /usr/local/src (or in your $HOME), and establish a symlink (ln -s) between /usr/local/bin/coot and the bin/coot of the freshly unpacked distribution.
If you then run coot, and the loader complains that a certain library is missing, just ask
yum whatprovides <thatlibrary>
and install the library, again using yum (assuming yum is available in your distribution, otherwise use apt or whatever is there for this purpose).
Installation on Debian/Ubuntu from debian archive files
As an alternative, you might wish to exploit the debain dpkg/apt/synaptic package management system on Ubuntu and other Debian linux distributions using (unofficial) pre-compiled debian packages for coot and its dependencies. (These will be discontinued when official packages become available.)
To do so, simply download the deb files and install with the command
sudo dpkg -i *.deb
dpkg will complain if a dependency is missing rather than install a broken program.
Converting to rpm packages
You can convert any debian package file into an rpm file using the program alien. I have done this and have made the resulting packages available.
Installation from source code via autobuild scripts
Installation of coot and all of its dependencies are handled automatically through the autobuild scripts. There are two versions:
- GTK1 - the old user interface. This script builds coot and all its dependencies.
- GTK2 - the new user interface. This script builds coot and most of the dependencies, excluding GTK2.
To build Coot, all you should need to do is edit a few settings in the top of the build script, or alternatively specify those settings as environment variables. For example, the following sequence of instructions will build the latest pre-release of the GTK 2 version with python support:
wget http://www.ysbl.york.ac.uk/~emsley/build-logs/build-it-gtk2-simple export AUTOBUILD_INSTALLED=${HOME}/autobuild/coot export AUTOBUILD_BUILD=${HOME}/autobuild/ export LOGS=$AUTOBUILD_BUILD/logs export NIGHTLY_DEST_DIR=$AUTOBUILD_BUILD export STABLE_DEST_DIR=$AUTOBUILD_BUILD export build_coot_prerelease=1 bash build-it-gtk2-simple python > build.log
(This script works in bash. For tcsh, replace 'export' with 'setenv' and '=' with ' '.
In some cases you may need to download additional development packages in order to build all the components.
Installation from source code manually
There are also instructions for Custom building Coot from source code.
Running Coot
General Topics
Controls
Stereographic Display
Side-by-Side
Hardware Stereo
Zalman Stereo
External Links
On-line User Manual
Coot's home page
Current mailing list archives
Mailing list archives: (no longer) current
Mailing list archives: 2004-05
Scheme Scripts
Coot can be scripted in scheme (guile) or python - support for each is more or less equal these days.
Several examples of coot extensions to the language can be seen by examining the 0-coot-state.scm file that coot leaves behind when it finishes.
COOPS
Coops generates a coot script from the output of molprobity, specifically probe, reduce, cluster and clashlist.
For an explanation of the principals underlying reduce and clashlist see the Dots Page. Get Molprobity software here.
Use Coot version 0.1 or higher.
Invoke like this (from the directory in which you run coot):
$ coops myfile.pdb
The use Calculate->Scripting to read in and run coops.scm
Get COOPS here.
Example Scheme Script 1: Move to Molecule Centres
This example can be found in the coot scheme sources (the function name is molecule-centres-gui and is in the xxx/share/coot/scheme/coot-gui.scm file). It is a simple function that creates a button box - a button for each coordinates molecule in Coot. It is annotated. Reproduced as coot-scheme1.
Example Scheme Script 2: Demo a Few of Coot's Features
Reading in a pdb file, an MTZ file and manipulating the model
This is a composite script and demonstrate reading pdb file, an MTZ file, translations, zoom, spin zooms, contour level changing, map masking, real space refinement, water addition and loop fitting.
The data files used in the example can be obtained here and here. Put them in the directory where you start coot. Save the script to your disk, then use Calculate -> Run Script... to activate it.
Example Scheme Script 3: Read CNS data
This CNS data reading script is a Cootenization of the CN2COOT script written by Joel Bard (it is based on his csh script) and can be used to compare and contrast scheme programming and shell script programming (the coot version is longer to some extent because it does extra error checking).
As well as doing the conversion the resulting mtz/maps are loaded into Coot.
It is part of Coot as of version 0.1.2.
Example Scheme Script 4: Load the Latest Data and PDB files Automatically
To load the most recent files, do this:
coot --script latest-files.scm
which enables the scripting function: (load-latest-files)
For extra gui goodness (you will need 0.1.2):
coot --script latest-files.scm --script extensions.scm
Example Scheme Script 5: Saving a Partial model
Here we create a small function to save part of a molecule and add a gui interface, it can be used in the usual way (i.e. with --script on the command line, Calculate->Run Script... or add the script to your ~/.coot file.
save-partial.scm
Example Scheme Script 6: Creating an interface for the Powermate Dial
The Powermate dial can be used with coot. One could just assign the rotations to +/-y keys and be done with it, but this script gives you a way of having positive and negative rotations in all three cartesian directions. The F1 key is mapped to positive rotation, the F2 key to negative rotation, and the F3 key permits you to toggle through x, y, and z, on successive key presses. I then map F1 and F2 into the ordinary rotations on the powermate (using send key equivalents) and then I map F3 into the single click on the dial, making it easy to toggle through x, y and z. The press-and-rotate options remain available; I map these into scroll up and down, and put them on the slowest response setting, which makes contouring density easier to control than it is from my mouse scroll wheel.
powermate-coot.scm
Example Scheme Script 7: Applying arbitrary value to "B" factor column
Imagine you have a file of some property (Chemical Shifts, for example) of a residue that you wish to apply to the atoms of a particular model from a pdb file as pseudo B factors. Here's how to do that in Coot:
We have a file "cs.tab" like this, the residue number then the chemical shift value (one for each residue in a particular chain):
1 1.53159
2 4.35884
3 4.07123
4 4.16932
5 6.69103
6 7.12071
7 10.7419
8 9.57176
Use apply-cs.scm to apply these values as pseudo temperature factors. Typical usage, where "A" is the chain id, and cs.tab the file of values per residue.
(apply-cs (read-pdb "test.pdb") "A" "cs.tab")
--script apply-cs.scm
Example Script 8: Partial Occupancy Dialog
Imagine that you have a structure that has residues with partial occupancy. After refinement, it would be convenient to quickly navigate to all such residues. How can that be done?
Start coot with command line arguments:
--script partial-occupancy-navigation.scm
This will provide an extra menu item called "Extras", clicking on "Residues with low occupancy..." therein will lead you through the process. Note that this will often work with SHELXL molecules, because they have atoms with negative (e.g -31) occupancies.
Note also that you will need a recent version of Coot to use this, as it stands. This will not work on stock Coot version 0.4.x. You can enable this for use with 0.4.x if you update/replace your xxx/share/coot/scheme/coot-gui.scm file from here.
Example Script 9: A GUI for Chopping Back Sidechains from a Residue Range
This is a simple interface to the delete-sidechain-range function, it illustrates how arguments can be transfered from the GUI to the scripting function. It was written in response to a question from Byron DeLaBarre.
Unfortunately (prior to 0.5) there was an error in the standard delete-sidechain-range function, which is why we over-ride it.
--script chop-side-chains-gui.scm
Example 10: How do I bind a key to Toggle the display of NCS ghosts?
With this script: toggle-ncs-ghosts-script
Example 11: Paul Emsley's Key Bindings
Just so you get an idea of the customization by key bindings here are what Paul uses currently (add to your .coot file).
Optional Wrappers and (External) Shell Script Enhancements
I (wgscott) wrote a coot wrapper shell script that lets you convert xplor/cns maps on the fly (you need to install mapman first) and has a few other enhancements.
I also made a Coot OS X applet that allows you to drag and drop a cns/xplor or ccp4 mapfile or any other coot-compatable file (mtz or pdb file, for example). Using the File > Get Info dialog, you can program this applet to open all .map and all .mtz files, if you want to, making these files double-clickable.
Download the Applet (requires a separate working coot installation)
Python Scripts
Example 1: Bernhard Lohkamp's Key Bindings
Just so you get an idea of the customization by key bindings here are what Bernhard/Paul uses currently (add to your .coot file or put the file in .coot-preferences directory).
bernhards_key_bindings_for_coot.py
Example 2: More key bindings (inspired by the Coot BB)
For (re-)colouring maps blue:
# bluefy last map
def blue_map_func1():
ls = map_molecule_list()
ls.reverse()
for map_mol in ls:
if (not map_is_difference_map(map_mol)):
set_map_colour(map_mol, 0.1, 0.5, 0.68)
graphics_draw()
break
add_key_binding("Blueify the Latest 2FoFc Map", "b",
lambda: blue_map_func1())
# bluefy all 2FoFc maps
def blue_map_func2():
for map_mol in map_molecule_list():
if (not map_is_difference_map(map_mol)):
set_map_colour(map_mol, 0.1, 0.5, 0.68)
graphics_draw()
add_key_binding("Blueify all 2FoFc Maps", "B",
lambda: blue_map_func2())
To (re-)colour coordinate molecules yellow:
# yellowify last coordinate molecule
# Note: this may overwrite the 'usual' short-cut of 'y' to add a terminal residue
def yellow_coords_func1():
ls = model_molecule_list()
if (len(ls) > 0):
set_molecule_bonds_colour_map_rotation(ls[-1], 20)
graphics_draw()
add_key_binding("Yellowify last coordinates", "y",
lambda: yellow_coords_func1())
# yellowify all coordinate molecules
add_key_binding("Yellowify all coordinates", "Y",
lambda: (map(lambda imol: set_molecule_bonds_colour_map_rotation(imol, 20),
model_molecule_list()),
graphics_draw()))
Python to Scheme and return
Translating between Python and Scheme
Python scripting is different to (default) scheme scripting which is mainly described in Paul Emsley's documentation (although it's mentioned somewhere, fairly hidden). You have to change the commands in the following way:
GUILE scripting: (guile-command argument1 argument2 ...)
PYTHON scripting: python_command(argument1, argument2, ...)
Simple rules for Scheme to Python translations
Here some simple rules how to translate from Scheme to Python. To translate the other way around, i.e. Python to Scheme, just turn the rules around:
- replace all '-' with '_' (except in equation when you need arithmetic '-' minus signs)
- move the brackets around the argument(s)
- separate multiple arguments by commas rather than spaces
- replace 'define' for functions with 'def' and for assignments with an '='
- Make sure to use indentation for the function content [Python is indentation sensitive] and a ':' after the function definition.
Some additional/advanced(?) rules:
- #f -> False
- #t -> True
- (set! variable value) -> variable=value
A simple example
In Scheme we may have the following script:
(define mol2-pdbFile "somePDBfile.pdb" )
(define mol2-model (read-pdb mol2-pdbFile))
(define (read-mol-again)
(clear-and-update-model-molecule-from-file mol2-model mol2-pdbFile))
(read-mol-again)
Which will translate into Python:
mol2_pdbFile = "somePDBfile.pdb"
mol2_model = read_pdb(mol2_pdbFile)
def read_mol_again():
clear_and_update_model_molecule_from_file(mol2_model, mol2_pdbFile)
read_mol_again()
Running a Scheme/Python command from Python/Scheme
As of Coot 0.5 (and if you have both scripting languages available) you an use the following commands to run a script or command in the other language:
(run-python-command "python_command(arg1, arg2, ...)") [from guile/scheme]
run_scheme_command("(scheme-command arg1 arg2 ...)") [from python]
Enhanced Menu Appearance
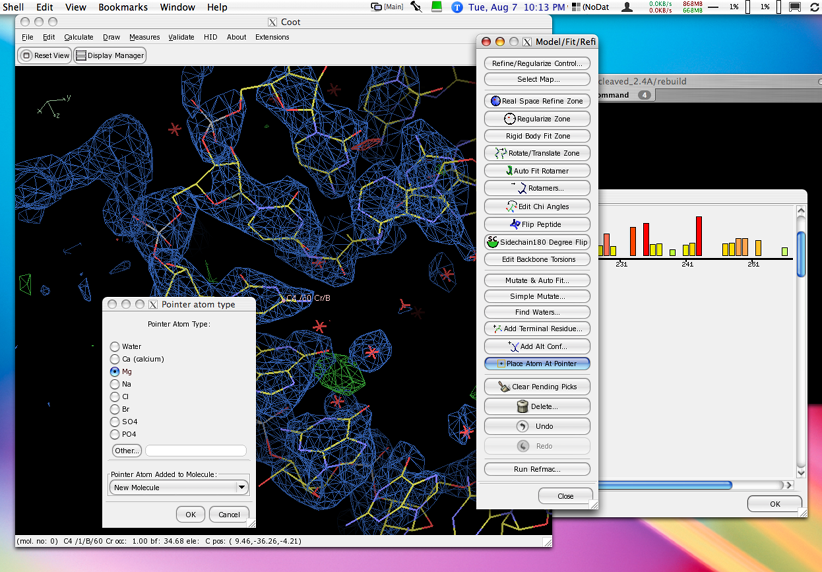
As of 0.4, coot works with gtk+2
This permits use of themes for a more OSX-like experience, among other things.
Click on the thumbnail image below to see a full-size screenshot of Coot with a gtk+2 Aqua-like theme.
To get this effect, you need the Glossy_P gtk+2 theme:
http://art.gnome.org/download/themes/gtk2/571/GTK2-Glossy_P.tar.gz
Edit a file called ~/.gtkrc-2.0 and put into it the following line:
include "/usr/share/themes/Glossy\ P/gtk-2.0/gtkrc"
Alternatively, if you use gnome or xfce4, you can open the theme manager and just make it open the downloaded Glossy_P tarball, and it should add this as a theme.
How can I get involved with Coot development?
Join the Coot Janitors project. This is a project to get new people involved in improving Coot, by acting as a clearing house for simple tasks which need doing, and providing documentation for doing them.
Assorted questions and answers (from the mailinglist)
It should be noted that the answers ("A") are from Paul Emsley himself (and were maybe slightly edited).
NCS edits
Q: I am sure this exists somewhere through scripting in COOT, but can I apply NCS edits to only a subset of NCS copies? In other words, can I tell coot which are NCS related chains, and which aren't. I am working on this nightmarish case of asymmetrical homodimers, where the sequences are very similar, but the structures are not, so I need to tell coot which chains are actually related to each other.
A: Nightmare. If you have a recent [1632 or later for the scheme version, 1646 for the python version] Coot, you can do this:
(manual-ncs-ghosts imol resno-start resno-end chain-id-list)
where
- imol is 0 (say)
- resno-start and resno-end is the residue range for the LSQ fitting to return the NCS matrix,
- chain-id-list is the list of chain-ids, starting with the master/reference chain-id and followed by the peer chain-ids that are NCS related, e.g. (list "A" "B" "D")
The python interface is similar.
There is also a GUI to activate this feature under Extensions -> NCS.
SHELXL
Description of problematic situation: I am using SHELXL to refine my 1.2 Å data and I am refining the hydrogen atoms. Subsequent rebuilding in coot is difficult though since hydrogens often does not "follow" when you do side chain rebuilding. For the moment I have quit transfering hydrogens to coot and add the hydrogens every refinement cycle, though it would be good I think if I could see them in coot without bothering about wrong positions. So these are my specific questions:
Specific Q1: Using "edit chi angles" does not work properly.
A: This fails because for chi angles Coot uses the Refmac dictionary to know what is connected to what (if it can). The work-around is to rename the refmac dictionary so Coot can't find it - which will force Coot to find bond by distance criteria.
Specific Q2: Using "real space refine" does not work properly.
A: Yes this fails. Hydrogens are named differently to SHELX hydrogens. In principal this could be made to work if the dictionary was reworked to use SHELX hydrogen names. This would also fix the chi angles problem too of course.
Image quality on NVidia cards
Q: improvement of image quality on machines with NVidia cards?
A: After talking about antialiasing with Stuart McNicholas, I discovered a program called nvidia-settings. I found that if I do:
Antialiasing Settings -> Override Application Setting, and set the slider to 16x
then I start Coot, I see that it starts up in nice antialiasing mode (which is a lot better that the poor mode that Coot has built-in). I don't know if this works on other/newer systems, but it works for me on my oldish GeForce 6600. (Of course the FPS takes a hit.)
Q: How to set the default to display symmetry related molecules?
A: Add (set-show-symmetry-master 1) to the appropriate file.
Startup files
Q: I still have a ".coot" file in my home folder for a few coot preferences that I couldn't find in the new ".coot-preferences/coot-preferences.scm". There is a warning that I should not add commands to this file. So is a "~/.coot" still the proper place to add default commands for coot?
A: Coot does not create a ~/.coot file for you, but will read it if it exists. Likewise, ~/.coot.py in which you can write python commands.~/.coot-preferences is a directory in which all .scm files and .py files are executed. coot-preferences.scm and coot_preferences.py there are generated by using the Edit -> Preferences dialog (and thus it overwrites older versions - hence the warning).
If you want to create a script that will be read by everyone then put those files into a directory defined by environment variable $COOT_SCHEME_EXTRAS_DIR (for *.scm) or $COOT_PYTHON_EXTRAS_DIR (*.py). All *.scm in $COOT_SCHEME_EXTRAS_DIR and *.py files in $COOT_PYTHON_EXTRAS_DIR will be executed at start up.
So you have a variety of places. Personally I mostly use ~/.coot.
Torsion general
Q: How do I use "torsion general"?
A: Thanks for pointing out the lack of documentation on this. I'll make a note to add some.
You need to click on the torsion-general icon, then click 4 atoms that describe the torsion - the first atom will be the base (non moving) part of the atom tree, on clicking the 4th atom a dialog will pop up with a "Reverse" button [1].
Move this dialog out of the way and then left mouse click and drag in the main window will rotate the moving/"top" part of the residue round the clicked atoms 2 and 3. When you are happy, click "Accept".
Window focus may be an issue - depending on your setting, the window manager may eat one of your clicks as you change focus between the dialog and the main graphics window (this I find annoying and there are instructions in the FAQ on how to turn that off for various systems).
[1] which may not work in 0.6-pre (grumble/sigh/sorry). If it doesn't not work, the "Reverse" button should invert the moving and "base" part of the residue.
Peak heights in maps
Q: I have some peaks in my map which take water or sodium/magnesium or chlorine atom with out giving out any positive or negative density upon further refinement. Is there any easy way of calculating the peak height / number of electrons at a given position, say a mouse click point in coot? Is there any formula to calculate the number of electrons based on sigma level and peak height, as given in difference map peaks in coot?
A: First, go to the Coot wiki and pick up the scheme key bindings.
If you want density information at a given cursor point: point at the blob, press the 'g' key (which recentres on the biggest density under the cursor).
using the Scheme scripting window:
(apply density-at-point (imol-refinement-map) (rotation-centre))
There is no user access to the peak integration code of coot as yet.